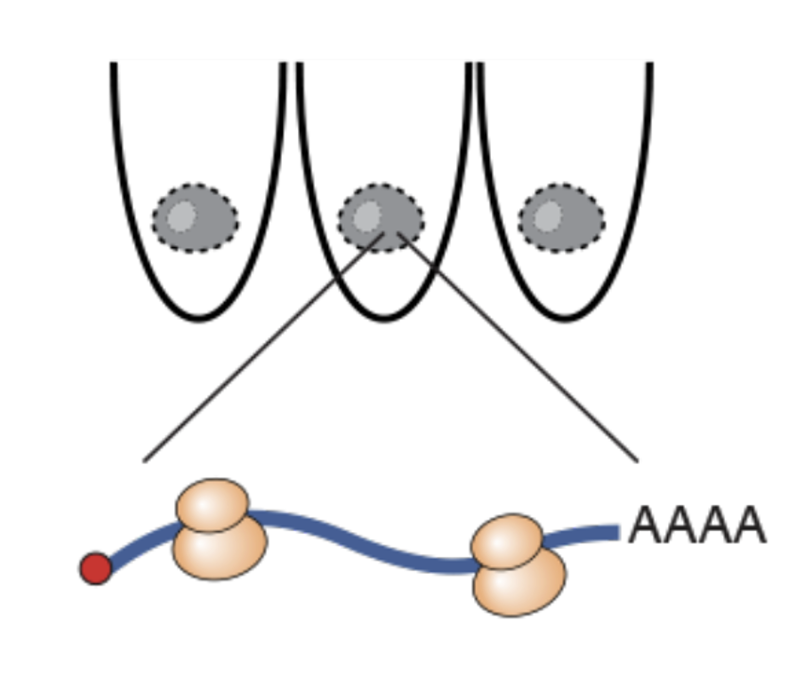
Mapping ribosomes in single cells
In recent years novel single-cell sequencing methods have allowed an in-depth analysis of the diversity of cell types and states in a wide range of organisms. Due to the continuous optimization of experimental and computational methods by many research groups, it is now possible to sequence the transcriptomes of thousands to millions of individual cells. Albeit an exciting development, transcription only covers the first step in the central dogma. The second step, the process of translation, is currently much harder to explore in single cells. Despite recent progress in detecting proteins by mass spectrometry with single-cell resolution it remains a major challenge to measure translation in individual cells. Building upon existing ribosome profiling protocols our laboratory recently majorly increased the sensitivity of these assays allowing ribosome profiling in single cells Integrated with a machine learning approach, this method achieves single-codon resolution in individual cells.
The major goal of this proposal is to further improve scRibo-seq to include a freezing and/or fixation step. Currently it is necessary to perform the scRibo-seq protocol immediately after sorting life cells, which severely hinders the processing of clinical material or material from non-local collaborating laboratories for which there is a delay between the harvesting of cells and the start of the scRibo-seq protocol. We will explore a range of different fixation protocols to study if these procedures are compatible with scRibo-seq. Next, we will use these protocols to simultaneously detect ribosome profiles and the abundance of transfer RNAs in single cells.